| CMap Home | Maps | Map Search | Feature Search | Matrix | Map Sets | Feature Types | Map Types | Evidence Types | Species | Saved Links | Help | Tutorial |
 |
|
*Bookmarks for this page will fail after this session expires.
Use the "Save Link" button to create a permanent link
|
| Map Type: | Genetic [ View Map Type Info ] | ||||||
|---|---|---|---|---|---|---|---|
| Map Set Name: | Cocoa - (Sca6xH)xC1 [ View Map Set Info ] | ||||||
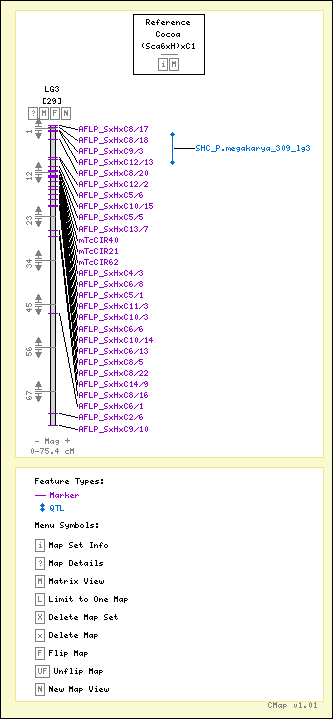
| Map Name: | LG3 | ||||||
| Map Start: | 0 cM | ||||||
| Map Stop: | 75.4 cM | ||||||
| Features by Type: |
|
| [ Download Map Data ]
[ Download Feature Correspondence Data ] |
||||||||
| Cocoa - (Sca6xH)xC1 - LG3 (Click headers to resort) |
Comparative Maps | |||||||
|---|---|---|---|---|---|---|---|---|
| Feature | Type | Position (cM) | Map | Feature | Type | Position | Evidence | Actions |
| AFLP_SxHxC9/10 | Marker | 75.4 | No other positions | |||||
| AFLP_SxHxC2/6 | Marker | 72.6 | No other positions | |||||
| AFLP_SxHxC6/1 | Marker | 47.5 | No other positions | |||||
| AFLP_SxHxC8/16 | Marker | 28 | No other positions | |||||
| AFLP_SxHxC14/9 | Marker | 26.4 | No other positions | |||||
| AFLP_SxHxC8/22 | Marker | 20.4 | No other positions | |||||
| AFLP_SxHxC8/5 | Marker | 20.3 | No other positions | |||||
| AFLP_SxHxC6/13 | Marker | 18.7 | No other positions | |||||
| AFLP_SxHxC10/14 | Marker | 17.5 | No other positions | |||||
| AFLP_SxHxC6/6 | Marker | 16.1 | No other positions | |||||
| AFLP_SxHxC10/3 | Marker | 15.5 | No other positions | |||||
| AFLP_SxHxC11/3 | Marker | 14.3 | No other positions | |||||
| AFLP_SxHxC5/1 | Marker | 14.1 | No other positions | |||||
| AFLP_SxHxC6/8 | Marker | 13.2 | No other positions | |||||
| AFLP_SxHxC4/3 | Marker | 12.9 | No other positions | |||||
| mTcCIR62 | Marker | 12.6 | Cocoa - TSH516 - LG3 | mTcCIR62 | Marker | 68 (cM) | Automated name-based | View Maps |
| Cocoa - consensus_disease_res - LG3 | mTcCIR62 | Marker | 16.7 (cM) | Automated name-based | View Maps | |||
| Cocoa - test - LG3 | mTcCIR62 | Marker | 16.7 (cM) | Automated name-based | View Maps | |||
| AFLP_SxHxC13/7 | Marker | 11.6 | No other positions | |||||
| AFLP_SxHxC5/5 | Marker | 11.4 | No other positions | |||||
| AFLP_SxHxC10/15 | Marker | 10.4 | No other positions | |||||
| AFLP_SxHxC5/6 | Marker | 8.7 | No other positions | |||||
| AFLP_SxHxC12/2 | Marker | 8.1 | No other positions | |||||
| AFLP_SxHxC8/20 | Marker | 3.7 | No other positions | |||||
| AFLP_SxHxC12/13 | Marker | 1.7 | No other positions | |||||
| AFLP_SxHxC9/3 | Marker | 1.4 | No other positions | |||||
| AFLP_SxHxC8/18 | Marker | 0.7 | No other positions | |||||